Hi everybody,
I have an edge list object weighted by a correlation value. I would like to know the changes in the node's connectivity on a continuous scale. That is how the node connectivity differs from the edge weight scale (-1) to the edge weight(+1). I would appreciate an idea of how to do it and what R package or statistical model could solve the problems etc.
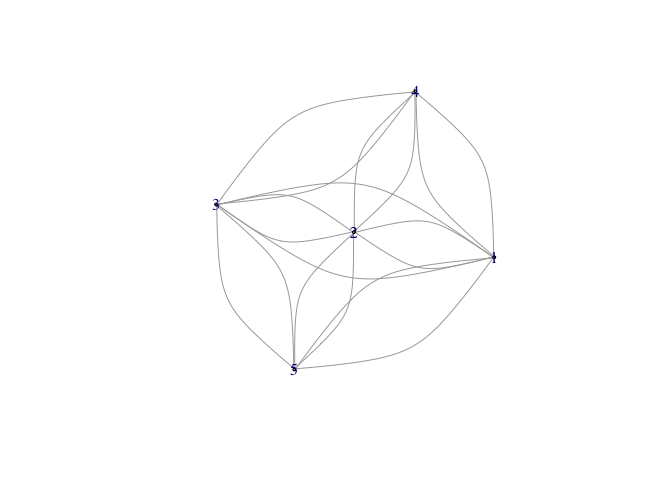
Look at the edge list object in the code below.
data <- structure(list(from = c(5L, 5L, 5L, 1L, 1L, 1L, 1L, 4L, 4L, 4L,
2L, 2L, 2L, 2L, 3L, 3L, 3L, 3L),
to = c(1L, 2L, 3L, 5L, 4L, 2L, 3L, 1L, 2L, 3L, 5L, 1L, 4L, 3L, 5L, 1L, 4L, 2L),
weight = c(runif(18, -1,1))),
row.names = c(NA, -18L),
class = "data.frame")
I appreciate your help!!
Best,
Amare