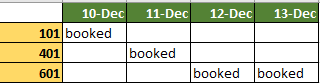
There's an idiom to error messages in R that takes a while to get attuned to. In this case it is directing attention to a "condition" in if(). The parameter in if() cond. From help(if)
A length-one logical vector that is not NA. Other types are coerced to logical if possible, ignoring any class. (As from R 4.2.0, conditions of length greater than one are an error.)
Translation: Take an object named diff and evaluate
length(diff)
If the return value is not 1 (it won't be here), the condition needs to be modified so that it is one. diff is created in the .local environment of the inner for loop based on the numeric value of the difference between two variables of the data object. To see what that does, evaluate it in the .Global environment.
dat <- data.frame(
Room = c(101, 401, 601),
Arrival = c("2019-12-10", "2019-12-11", "2019-12-12"),
Departure = c("2019-12-11", "2019-12-13", "2019-12-15"))
dat |> str()
#> 'data.frame': 3 obs. of 3 variables:
#> $ Room : num 101 401 601
#> $ Arrival : chr "2019-12-10" "2019-12-11" "2019-12-12"
#> $ Departure: chr "2019-12-11" "2019-12-13" "2019-12-15"
# convert character reprsentation of dates to date objects
# since it will come up frequently, make it an object
datify <- function(x,y) lubridate::ymd(x[y][[1]])
dat[2] <- datify(dat,2)
dat[3] <- datify(dat,3)
dat |> str()
#> 'data.frame': 3 obs. of 3 variables:
#> $ Room : num 101 401 601
#> $ Arrival : Date, format: "2019-12-10" "2019-12-11" ...
#> $ Departure: Date, format: "2019-12-11" "2019-12-13" ...
(length(diff <- (dat[3] - dat[2])[[1]])) == 1
#> [1] FALSE
# create function, just for practice
is_length0 <- function(x,y,z) {
x[y] = datify(x,y)
x[z] = datify(x,z)
(length(diff <- (x[z] - x[y])[[1]])) == 1
}
is_length0(dat,2,3)
#> [1] FALSE
Created on 2023-01-27 with reprex v2.0.2
Note that the effect of wrapping an expression in () to immediately evaluate it.
(Now is a good time to get in the habit of avoiding built-in functions as user-created objects—data is one, and df another. It's often possible to get away with it, but sooner or later some operation is going to give priority in namespace to the function object and complain when it feels that it is being mistreated as a data object and there may be an error message to the effect that can't subset a closure.)
for loops are ok in R and sometimes convenient, with important conditions:
- Unlike
C and its progeny, what happens in a for loop stays there until it is returned.
- It is clearer, and faster when dealing with moderately large objects to pre-allocate a receiver object outside the loop
holder <- vector(length = 1e5)
for(i in seq_along(something) holder[i] = some_function(some_arguments
- vectorized equivalents exist and can be faster
apply(mtcars,1,mean)
- Involved control statements can be written in
Python or C/C++ and called in an R script through the {reticulate} or {Rcpp} packages.
Finally, it is profitable to think of R in terms of its original intent and continuing strength. It presents to the user as a functional rather than a procedural language using the school algebra paradigm of f(x) = y.
x is what is to hand, y is what is desired and f is a function that will transform the one into the other. Any of these objects (in R everything is an object, even functions) can be composite in the tradition of f(g(x) = y. The virtue of this approach is the focus it requires on what the objects are and do, rather than the procedural/imperative process of focusing on how to express the transformation in a stepwise manner.