Hi all
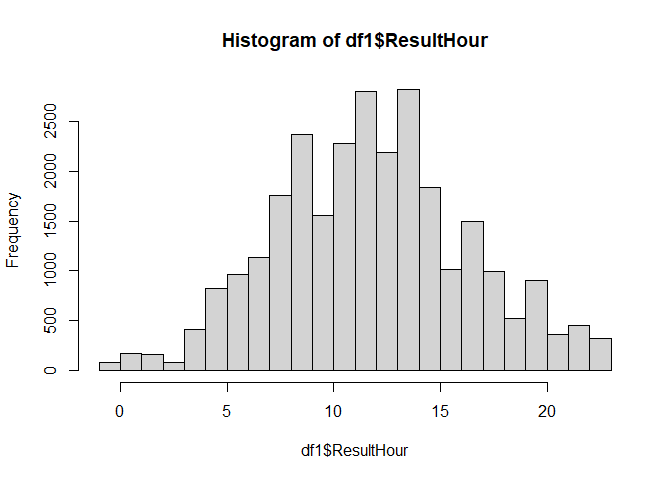
I have a dataset that contains testsamples and there result time. I want to visualize the amount of samples each hour and visualize it on a histogram. I use following code:
#Calculate the amount of Result-samples each hour:
df2 <- as.data.frame(hour(hms(df1_filtered$ResultTime)))
df_aggr_Result <- aggregate(df2, by=list(df2$`hour(hms(df1_filtered$ResultTime))`), FUN = length)
When a test is performed each hour, in the first column of the df_aggr_Result, the hours are going from 0 until 23 (like it supposed to be). I have a good histogram.
The problem occurs when eg. a test is done 7 times at 12h and 3 times at 18h. When I use the code above, I get a df_aggr_Result with only 2 rows (one row with 12h and the other with 18h), this results in a histogram with two "bars" and not the wanted 24 bars.
How can I code that the first column of the created dataframe counts from 0 untill 23 and add a 0 in the other column if it cannot be counted in the previous dataframe.
Here is a link to my intial CSV:
https://drive.google.com/file/d/14KSe_pdATXQJTZSJMOHKWExA_m7XCgJH/view?usp=sharing
Thanks in advance!
This is my total code:
knitr::opts_chunk$set(echo = TRUE)
install.packages(c("lubridate"),repos = "http://cran.us.r-project.org")
install.packages(c("dplyr"),repos = "http://cran.us.r-project.org")
knitr::include_graphics("https://weareofficemanagementmechelen.files.wordpress.com/2019/12/roche-logo.jpeg?w=400")
#First the exported file from the Infinity server needs to be in the correct folder and read in:
setwd("G:/My Drive/Traineeship Advanced Bachelor of Bioinformatics 2022/Internship 2022-2023/internship documents/")
server_data <- read.delim(file="PostCheckAnalysisRoche.txt", header = TRUE, na.strings=c(""," ","NA"))
#Create a subset of the data to remove/exclude the unnecessary columns:
subset_data_server <- server_data[,c(-5:-7,-9,-17,-25:-31)]
#Remove the rows with blank/NA values:
df1 <- na.omit(subset_data_server)
#The difference between the ResultTime and the FirstScanTime is the turn around time:
df1$TS_start <- paste(df1$FirstScanDate, df1$FirstScanTime)
df1$TS_end <- paste(df1$ResultDate, df1$ResultTime)
df1$TAT <- difftime(df1$TS_end,df1$TS_start,units = "mins")
library(lubridate)
library(dplyr)
selectInput(inputId='test', label='Test:',
choices = df1$TestName)
df1_filtered <- df1[df1$TestName == "IGF",]
#Calculate the amount of Result-samples each hour:
df2 <- as.data.frame(hour(hms(df1_filtered$ResultTime)))
df_aggr_Result <- aggregate(df2, by=list(df2$`hour(hms(df1_filtered$ResultTime))`), FUN = length)
#Renaming
names(df_aggr_Result)[names(df_aggr_Result) == "Group.1"] <- "hour"
names(df_aggr_Result)[names(df_aggr_Result) == "hour(hms(df1_filtered$ResultTime))"] <- "amount of samples"
renderPlot({
x <- as.numeric(df_aggr_Result$`amount of samples`)
hist(x, breaks = 24,
xlab = "Hours in a day",
ylab = "Amount of samples"
)
})