Yes, dots are essentially used for two things: first, to pass arguments to other functions, second, to accept arbitrary arguments.
Here is an example of the first:
mean_of_half <- function(x, ...){
half <- x/2
mean(half, ...)
}
mean_of_half(c(1, 10, NA), na.rm = TRUE)
#> [1] 2.75
Created on 2022-11-06 by the reprex package (v2.0.1)
In this case, mean_of_half() is a function which just calls the R builtin mean(), after dividing its input. What is great with the ... is that since I pass them to mean(), when using mean_of_half() I can give any argument that mean() understands, and I don't need to define them myself when writing mean_of_half(). I don't even need to know they exist.
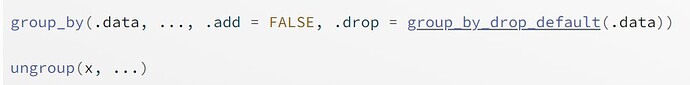
An example of the second case is dplyr::group_by(). In the source code of group_by(), there is no assumption about what the possible column names are, it can accept anything. Here you are not "passing the dots" to another function, you are using them directly. A very simple example could be:
print_dots <- function(...){
list(...)
}
print_dots(a=5, b=7)
#> $a
#> [1] 5
#>
#> $b
#> [1] 7
Created on 2022-11-06 by the reprex package (v2.0.1)
Sure, that would be a solution. But how can you ensure it? First, there are two "you" in that sentence: the person writing the function, and the person using it.
Technically, it's on the user to read the manual and ensure that the arguments they use are compatible with the function. But there are several problems: as a typical user, do you fully read the entire manual of each and every function you use? I know I don't, I will typically read the parts that I need, and I will rely on my memory for functions I used before. And what if new arguments get added? For example dplyr::mutate() got its arguments before and after somewhat recently. It's totally possible that I have data frames with columns of that name, that means my code that I had written and that used to work will suddenly break after an update. Oh, and what if you, the user, are combining two functions written by other people? If ... is being passed to an existing function you don't get to choose how its arguments are named.
So, when you're writing such a function, you'd rather assume that the user may be stupid and not read the documentation, and make sure that really there is (almost) no way for a user to accidentally use arguments with the same name in .... Which is why mutate() actually called these arguments .before and .after, because it's very unlikely that these are column names in my data frames.