Thanks for the reply and the link! I did check here but seemed to only be for parametric approach and also seemed to only be for linear models:
(Scrolling to bottom)
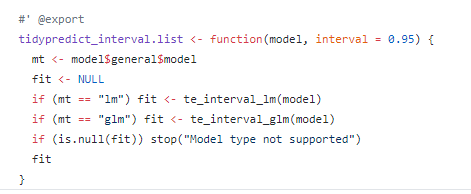
I would be interested in an example that uses simulation based approach as well as example for non-linear model.
