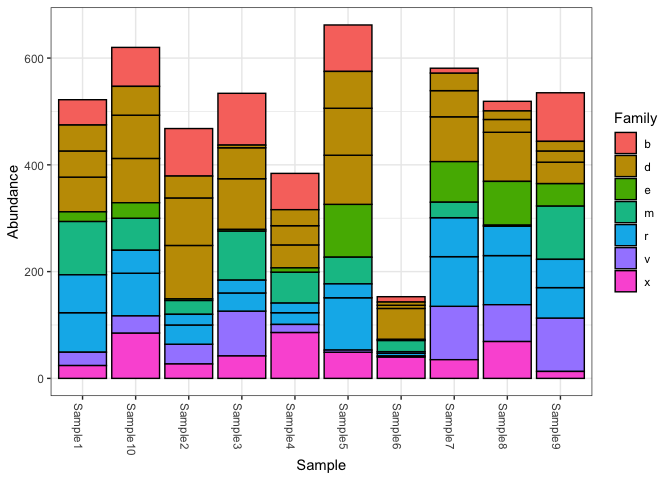
Currently, I am trying to draw a taxa bar plot using "phyloseq" and "ggplot2".
I assumed that the fct_reorder function can solve this problem. However, applying this function is difficult for me. ex) where should I use this function in my codes.
I should have used the "Reprex" package, however, it did not work with some errors. Therefore, I made my code a little messy.
Thank you for the expert's help.
ASVmat <-tibble::tribble(
~ASV, ~CON_D1_08611, ~CON_D1_10428, ~CON_D1_35264, ~TRT_D1_08620, ~TRT_D1_11441, ~TRT_D1_84032,
"ASV1", 25.46, 0.18, 5.01, 67.56, 16.73, 35.52,
"ASV2", 15.98, 5.81, 1.09, 0.07, 13.6, 0,
"ASV3", 13.24, 1.94, 0.04, 0.01, 0, 0,
"ASV4", 11.73, 0.01, 10.41, 0.29, 5, 1.03,
"ASV5", 8.9, 0.02, 16.68, 0, 0.01, 0.09,
"ASV6", 7.97, 0, 0.03, 0.53, 33.7, 0,
"ASV7", 6.22, 0, 0.03, 0, 0.02, 0,
"ASV8", 2.41, 6.76, 0.02, 0, 0.4, 0,
"ASV9", 1.83, 0, 0.03, 4.58, 0.16, 0.01,
"ASV10", 1.8, 0.79, 0, 0, 0, 0,
"ASV11", 0.72, 17.15, 0, 0, 0, 0,
"ASV12", 0.69, 10.37, 24.5, 1.76, 3.28, 0.14,
"ASV13", 0.64, 4.94, 0.02, 0, 0.34, 0,
"ASV14", 0.63, 2.3, 0.01, 0, 0, 0,
"ASV15", 0.28, 1.19, 0.05, 0.2, 0.1, 0,
"ASV16", 0.28, 0.35, 0.01, 0.01, 0, 0,
"ASV17", 0.21, 0.17, 0, 0, 14.08, 0,
"ASV18", 0.2, 0.09, 25.2, 17.18, 2.49, 2.67,
"ASV19", 0.19, 0.48, 0, 0, 0.87, 0,
"ASV20", 0.18, 0, 0.01, 0.01, 0, 0,
"ASV21", 0.14, 0.01, 0.14, 0.14, 0.2, 0,
"ASV22", 0.09, 9.53, 0.09, 0, 0, 0,
"ASV23", 0.04, 0, 0, 0.05, 0.78, 0,
"ASV24", 0.03, 0, 0.27, 0.13, 0, 0,
"ASV25", 0.02, 0, 0.2, 0.01, 0, 0,
"ASV26", 0.02, 0.01, 0.04, 0.01, 0.64, 0.01,
"ASV27", 0.01, 0, 2.54, 0.42, 0.04, 33.93,
"ASV28", 0.01, 0.03, 0, 0.03, 0.06, 0.26,
"ASV29", 0.01, 0.43, 0, 0.01, 0, 0,
"ASV30", 0.01, 15.67, 3.66, 0.91, 1.75, 1.05,
"ASV31", 0.01, 0, 1.84, 0.51, 0.03, 25.22,
"ASV32", 0.01, 0, 0.75, 4.16, 0, 0,
"ASV33", 0.01, 0.03, 0.13, 0.05, 0, 0,
"ASV34", 0.01, 0.01, 0.17, 0.28, 0, 0,
"ASV35", 0, 1.35, 0.07, 0.32, 0.23, 0.02,
"ASV36", 0, 0.02, 0.04, 0.01, 2.05, 0.03,
"ASV37", 0, 0, 0, 0, 0.18, 0.02,
"ASV38", 0, 0, 2.7, 0.01, 0, 0,
"ASV39", 0, 10.11, 0.19, 0, 0, 0,
"ASV40", 0, 8.33, 0, 0.01, 0, 0,
"ASV41", 0, 0.35, 3.73, 0, 0.98, 0,
"ASV42", 0, 0.47, 0.03, 0.03, 0, 0,
"ASV43", 0, 0.48, 0.01, 0, 0, 0,
"ASV44", 0, 0.31, 0.06, 0.04, 0.38, 0,
"ASV45", 0, 0.18, 0.01, 0, 1.02, 0,
"ASV46", 0, 0, 0.03, 0.11, 0.67, 0,
"ASV47", 0, 0, 0, 0.28, 0.01, 0,
"ASV48", 0, 0, 0, 0.15, 0.03, 0
)
TAXmat <- tibble::tribble(
~ASV, ~Kingdom, ~Phylum, ~Class, ~Order, ~Family, ~Genus,
"ASV1", "Bacteria", "Firmicutes", "Bacilli", "Lactobacillales", "Streptococcaceae", "Streptococcus",
"ASV2", "Bacteria", "Bacteroidota", "Bacteroidia", "Bacteroidales", "Bacteroidaceae", "Bacteroides",
"ASV3", "Bacteria", "Firmicutes", "Clostridia", "Oscillospirales", "Butyricicoccaceae", "Butyricicoccus",
"ASV4", "Bacteria", "Firmicutes", "Bacilli", "Lactobacillales", "Enterococcaceae", "Enterococcus",
"ASV5", "Bacteria", "Actinobacteriota", "Actinobacteria", "Bifidobacteriales", "Bifidobacteriaceae", "Bifidobacterium",
"ASV6", "Bacteria", "Verrucomicrobiota", "Verrucomicrobiae", "Verrucomicrobiales", "Akkermansiaceae", "Akkermansia",
"ASV7", "Bacteria", "Firmicutes", "Negativicutes", "Veillonellales-Selenomonadales", "Veillonellaceae", "Veillonella",
"ASV8", "Bacteria", "Firmicutes", "Clostridia", "Lachnospirales", "Lachnospiraceae", "[Ruminococcus]_gnavus_group",
"ASV9", "Bacteria", "Proteobacteria", "Gammaproteobacteria", "Enterobacterales", "Pasteurellaceae", "Gallibacterium",
"ASV10", "Bacteria", "Firmicutes", "Clostridia", "Oscillospirales", "Ruminococcaceae", "Fournierella",
"ASV11", "Bacteria", "Firmicutes", "Clostridia", "Oscillospirales", "Ruminococcaceae", "Faecalibacterium",
"ASV12", "Bacteria", "Firmicutes", "Clostridia", "Clostridiales", "Clostridiaceae", "Clostridium_sensu_stricto_1",
"ASV13", "Bacteria", "Actinobacteriota", "Coriobacteriia", "Coriobacteriales", "Coriobacteriaceae", "Collinsella",
"ASV14", "Bacteria", "Bacteroidota", "Bacteroidia", "Bacteroidales", "Prevotellaceae", "Alloprevotella",
"ASV15", "Bacteria", "Firmicutes", "Bacilli", "Erysipelotrichales", "Erysipelatoclostridiaceae", "Erysipelatoclostridium",
"ASV16", "Bacteria", "Firmicutes", "Clostridia", "Oscillospirales", "Ruminococcaceae", "Ruminococcus",
"ASV17", "Bacteria", "Fusobacteriota", "Fusobacteriia", "Fusobacteriales", "Fusobacteriaceae", "Fusobacterium",
"ASV18", "Bacteria", "Proteobacteria", "Gammaproteobacteria", "Enterobacterales", "Enterobacteriaceae", "Escherichia-Shigella",
"ASV19", "Bacteria", "Firmicutes", "Negativicutes", "Acidaminococcales", "Acidaminococcaceae", "Phascolarctobacterium",
"ASV20", "Bacteria", "Firmicutes", "Clostridia", "Lachnospirales", "Lachnospiraceae", "Howardella",
"ASV21", "Bacteria", "Firmicutes", "Clostridia", "Clostridiales", "Clostridiaceae", "Clostridium_sensu_stricto_2",
"ASV22", "Bacteria", "Firmicutes", "Clostridia", "Lachnospirales", "Lachnospiraceae", "Tyzzerella",
"ASV23", "Bacteria", "Actinobacteriota", "Coriobacteriia", "Coriobacteriales", "Eggerthellaceae", "Paraeggerthella",
"ASV24", "Bacteria", "Firmicutes", "Clostridia", "Oscillospirales", "Oscillospiraceae", "Oscillospiraceae_UCG-002",
"ASV25", "Bacteria", "Proteobacteria", "Gammaproteobacteria", "Burkholderiales", "Sutterellaceae", "Parasutterella",
"ASV26", "Bacteria", "Firmicutes", "Clostridia", "Eubacteriales", "Eubacteriaceae", "Eubacterium",
"ASV27", "Bacteria", "Firmicutes", "Bacilli", "Lactobacillales", "Lactobacillaceae", "Limosilactobacillus",
"ASV28", "Bacteria", "Proteobacteria", "Gammaproteobacteria", "Pseudomonadales", "Moraxellaceae", "Psychrobacter",
"ASV29", "Bacteria", "Firmicutes", "Bacilli", "Erysipelotrichales", "Erysipelotrichaceae", "Faecalicoccus",
"ASV30", "Bacteria", "Others", "Others", "Others", "Others", "Others",
"ASV31", "Bacteria", "Firmicutes", "Bacilli", "Lactobacillales", "Lactobacillaceae", "Ligilactobacillus",
"ASV32", "Bacteria", "Firmicutes", "Clostridia", "Peptostreptococcales-Tissierellales", "Peptostreptococcaceae", "Terrisporobacter",
"ASV33", "Bacteria", "Firmicutes", "Clostridia", "Christensenellales", "Christensenellaceae", "Christensenellaceae_R-7_group",
"ASV34", "Bacteria", "Firmicutes", "Clostridia", "Oscillospirales", "Oscillospiraceae", "Oscillospiraceae_NK4A214_group",
"ASV35", "Bacteria", "Firmicutes", "Clostridia", "Lachnospirales", "Lachnospiraceae", "Blautia",
"ASV36", "Bacteria", "Actinobacteriota", "Actinobacteria", "Corynebacteriales", "Corynebacteriaceae", "Corynebacterium",
"ASV37", "Bacteria", "Proteobacteria", "Gammaproteobacteria", "Burkholderiales", "Comamonadaceae", "Comamonas",
"ASV38", "Bacteria", "Proteobacteria", "Alphaproteobacteria", "Rhodospirillales", "Others", "Others",
"ASV39", "Bacteria", "Verrucomicrobiota", "Chlamydiae", "Chlamydiales", "Chlamydiaceae", "Chlamydia",
"ASV40", "Bacteria", "Firmicutes", "Clostridia", "Clostridia_UCG-014", "Clostridia_UCG-014", "Clostridia_UCG-014",
"ASV41", "Bacteria", "Bacteroidota", "Bacteroidia", "Bacteroidales", "Tannerellaceae", "Parabacteroides",
"ASV42", "Bacteria", "Firmicutes", "Clostridia", "Oscillospirales", "Oscillospiraceae", "Others",
"ASV43", "Bacteria", "Firmicutes", "Clostridia", "Oscillospirales", "Ruminococcaceae", "Others",
"ASV44", "Bacteria", "Firmicutes", "Clostridia", "Lachnospirales", "Lachnospiraceae", "Lachnoclostridium",
"ASV45", "Bacteria", "Proteobacteria", "Gammaproteobacteria", "Burkholderiales", "Sutterellaceae", "Sutterella",
"ASV46", "Bacteria", "Firmicutes", "Clostridia", "Peptostreptococcales-Tissierellales", "Peptostreptococcaceae", "Peptostreptococcus",
"ASV47", "Bacteria", "Proteobacteria", "Gammaproteobacteria", "Enterobacterales", "Morganellaceae", "Proteus",
"ASV48", "Bacteria", "Bacteroidota", "Bacteroidia", "Bacteroidales", "Prevotellaceae", "Prevotellaceae_UCG-003"
)
Samplemat <- tibble::tribble(
~Sample, ~Day, ~Group,
"CON_D1_08611", "D_1", "Healthy",
"CON_D1_10428", "D_1", "Healthy",
"CON_D1_35264", "D_1", "Healthy",
"TRT_D1_08620", "D_1", "Virus",
"TRT_D1_11441", "D_1", "Virus",
"TRT_D1_84032", "D_1", "Virus"
)
library(phyloseq)
library(ggplot2)
library(tidyverse)
library(readxl)
library(RColorBrewer)
library(reprex)
ASVmat = read_tsv("Metagenomic/Day1/ASVmat.tsv") %>%
tibble::column_to_rownames("ASV")
TAXmat = read_tsv("Metagenomic/Day1/TAXmat.tsv") %>%
tibble::column_to_rownames("ASV")
Samplemat = read_tsv("Metagenomic/Day1/Sample.tsv") %>%
tibble::column_to_rownames("Sample")
ASVmat <- as.matrix(ASVmat)
TAXmat <- as.matrix(TAXmat)
ASV = otu_table(ASVmat, taxa_are_rows = TRUE)
TAX = tax_table(TAXmat)
samples = sample_data(Samplemat)
carbom <- phyloseq(ASV, TAX, samples)
phylum_colors <- c("#1B9E77", "#D95F02", "#7570B3", "#E7298A", "#66A61E", "#E6AB02","#A6761D", "#666666",
"#1B9E77", "#D95F02", "#7570B3", "#E7298A", "#66A61E", "#E6AB02","#A6761D", "#666666",
"#1B9E77", "#D95F02", "#7570B3", "#E7298A", "#66A61E", "#E6AB02","#A6761D", "#666666",
"#1B9E77", "#D95F02", "#7570B3", "#E7298A", "#66A61E", "#E6AB02","#A6761D", "#666666",
"#1B9E77", "#D95F02", "#7570B3", "#E7298A", "#66A61E", "#E6AB02","#A6761D", "#666666",
"#1B9E77", "#D95F02", "#7570B3", "#E7298A", "#66A61E", "#E6AB02","#A6761D", "#666666",
"#1B9E77", "#D95F02", "#7570B3", "#E7298A", "#66A61E", "#E6AB02","#A6761D", "#666666",
"#1B9E77", "#D95F02", "#7570B3", "#E7298A", "#66A61E", "#E6AB02","#A6761D", "#666666")
Phylum_lists <- c("Firmicutes","Proteobacteria", "Bacteroidota","Actinobacteriota", "Verrucomicrobiota", "Fusobacteriota", "Others")
plot_bar(carbom, fill = "Phylum") +
geom_bar(aes(fill = Phylum), stat = "identity", position = "fill") +
facet_grid(~Group, scales = "free") +
scale_fill_brewer(palette = "Dark2") +
scale_fill_manual(values = phylum_colors) +
scale_y_continuous(name = "Relative abundance (%)") +
scale_y_continuous(expand=c(0, 0)) +
guides(fill = guide_legend(ncol=4)) +
theme_classic() +
theme(axis.text.x = element_blank(),
axis.text.y = element_text(color = "black"),
axis.title.x = element_blank(),
legend.position = "bottom",
legend.key.size = unit(0.6, 'cm'),
legend.key.height = unit(0.6, "cm"),
legend.title = element_blank(),
strip.text = element_text(face = "bold"),
strip.background = element_blank())
Rplot01.pdf (7.6 KB)