I have one response variable and 4 predictors and I am performing a spatial random forest regression at a coarse spatial scale. My goal is to take the model parameters and apply them to a finer spatial resolution in order to predict the response variable at the finer spatial scale.
When I run
p <- stats::predict(object = model.spatial, # name of the spatialRF model
data = s, # data.frame containing the predictors at the fine spatial scale (without NaN values)
type = "response")$predictions
I am getting this error: Error in predict.ranger.forest(forest, data, predict.all, num.trees, type,: Error: One or more independent variables not found in data.
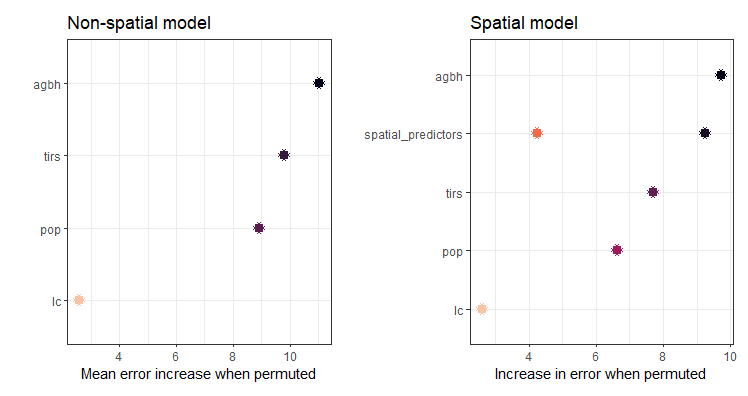
The reason I am getting this error is because when I run the spatial version of RF, one extra predictor is created:
Using the spatial random forest model, how can make predictions at the finer spatial scale, taking into account that now I have one more predictor variable?
Here is the code:
library(spatialRF)
library(stats)
wd = "path/"
block.data = read.csv(paste0(wd, "block.data.csv"))
#names of the response variable and the predictors
dependent.variable.name <- "ntl"
predictor.variable.names <- colnames(block.data)[4:7]
#coordinates of the cases
xy <- block.data[, c("x", "y")]
block.data$x <- NULL
block.data$y <- NULL
#distance matrix
distance.matrix <- as.matrix(dist(block.data))
min(distance.matrix)
max(distance.matrix)
#distance thresholds (same units as distance_matrix)
distance.thresholds <- c(0, 20, 50, 100, 200, 500)
#random seed for reproducibility
random.seed <- 456
#creating and registering the cluster
local.cluster <- parallel::makeCluster(
parallel::detectCores() - 1,
type = "PSOCK")
doParallel::registerDoParallel(cl = local.cluster)
# fitting a non-spatial Random Forest
model.non.spatial <- spatialRF::rf(
data = block.data,
dependent.variable.name = dependent.variable.name,
predictor.variable.names = predictor.variable.names,
distance.matrix = distance.matrix,
distance.thresholds = distance.thresholds,
xy = xy,
seed = random.seed,
verbose = FALSE)
# Fitting a spatial model with rf_spatial()
model.spatial <- spatialRF::rf_spatial(
model = model.non.spatial,
method = "mem.moran.sequential", #default method
verbose = FALSE,
seed = random.seed)
#stopping the cluster
parallel::stopCluster(cl = local.cluster)
# export residuals of the spatialRF model at the coarse scale
rsds = as.data.frame(cbind(xy, model.spatial$residuals$values))
colnames(rsds)[3] = "resids"
coordinates(rsds) <- ~ x + y
gridded(rsds) <- TRUE
rsds <- raster(rsds)
crs(rsds) = provoliko
writeRaster(rsds, paste0(wd, "srf_resids.tif"), overwtite = TRUE)
# prediction at a finer spatial scale
s = read.csv(paste0(wd, "s.csv"))
p <- stats::predict(object = model.spatial,
data = s,
type = "response")$predictions
You can download the a small sample of the data from here.